| Virus classification | |
|---|---|
| (unranked): | Virus |
| Realm: | Riboviria |
| Phylum: | incertae sedis |
| Order: | Nidovirales |
| Family: | Coronaviridae |
| Subfamily: | Orthocoronavirinae |
| Genera[1] | |
| Synonyms[2][3][4] | |
| |
Coronaviruses constitute the subfamilyOrthocoronavirinae, in the family Coronaviridae, order Nidovirales, and realm Riboviria.[5][6] They are enveloped viruses with a positive-sense single-stranded RNA genome and a nucleocapsid of helical symmetry. The genome size of coronaviruses ranges from approximately 27 to 34 kilobases, the largest among known RNA viruses.[7]The name coronavirus is derived from the Latin corona, meaning "crown" or "halo", which refers to the characteristic appearance reminiscent of a crown or a solar corona around the virions (virus particles) when viewed under two-dimensional transmission electron microscopy, due to the surface covering in club-shaped protein spikes.
Discovery
Coronaviruses were first discovered in the late 1960s.[8] The earliest ones discovered were an infectious bronchitis virus in chickens and two in human patients with the common cold (later named human coronavirus 229E and human coronavirus OC43).[9] Other members of this family have since been identified, including SARS-CoV in 2003, HCoV NL63 in 2004, HKU1 in 2005, MERS-CoV in 2012, and SARS-CoV-2(formerly known as 2019-nCoV) in 2019. Most of these have involved serious respiratory tract infections.
Etymology
The name "coronavirus" is derived from Latin corona, meaning "crown" or "wreath", itself a borrowing from Greekκορώνη korṓnē, "garland, wreath". The name refers to the characteristic appearance of virions (the infective form of the virus) by electron microscopy, which have a fringe of large, bulbous surface projections creating an image reminiscent of a crown or of a solar corona.[citation needed] This morphology is created by the viral spike peplomers, which are proteins on the surface of the virus.
Morphology
Coronaviruses are large pleomorphicspherical particles with bulbous surface projections.[10] The diameter of the virus particles is around 120 nm.[11] The envelope of the virus in electron micrographs appears as a distinct pair of electron dense shells.[12]
The viral envelope consists of a lipid bilayer where the membrane (M), envelope (E) and spike (S) structural proteins are anchored.[13] A subset of coronaviruses (specifically the members of Betacoronavirus subgroup A) also have a shorter spike-like surface protein called hemagglutinin esterase (HE).[5]
Inside the envelope, there is the nucleocapsid, which is formed from multiple copies of the nucleocapsid (N) protein, which are bound to the positive-sense single-stranded RNA genome in a continuous beads-on-a-string type conformation.[11][14] The genome sizefor coronaviruses ranges from approximately 27 to 34 kilobases.[7] The lipid bilayer envelope, membrane proteins, and nucleocapsid protect the virus when it is outside the host cell.[15]
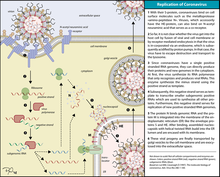
Replication
Infection begins when the virus entersthe host organism and the spike protein attaches to its complementary host cell receptor. After attachment, a proteaseof the host cell cleaves and activates the receptor-attached spike protein. Depending on the host cell protease available, cleavage and activation allows cell entry through endocytosis or direct fusion of the viral envelop with the host membrane.[16]
On entry into the host cell, the virus particle is uncoated, and its genomeenters the cell cytoplasm.[17] The coronavirus RNA genome has a 5′ methylated cap and a 3′ polyadenylated tail, which allows the RNA to attach to the host cell's ribosome for translation.[18] The host ribosome translates the initial overlapping open reading frame of the virus genome and forms a long polyprotein. The polyprotein has its own proteases which cleave the polyprotein into multiple nonstructural proteins.[19]
A number of the nonstructural proteins coalesce to form a multi-proteinreplicase-transcriptase complex (RTC). The main replicase-transcriptase protein is the RNA-dependent RNA polymerase(RdRp). It is directly involved in the replication and transcription of RNA from an RNA strand. The other nonstructural proteins in the complex assist in the replication and transcription process. The exoribonuclease non-structural protein for instance provides extra fidelity to replication by providing a proofreadingfunction which the RNA-dependent RNA polymerase lacks.[20]
One of the main functions of the complex is to replicate the viral genome. RdRp directly mediates the synthesis of negative-sense genomic RNA from the positive-sense genomic RNA. This is followed by the replication of positive-sense genomic RNA from the negative-sense genomic RNA.[19] The other important function of the complex is to transcribe the viral genome. RdRp directly mediates the synthesis of negative-sense subgenomic RNA molecules from the positive-sense genomic RNA. This is followed by the transcription of these negative-sense subgenomic RNA molecules to their corresponding positive-sense mRNAs.[19]
The replicated positive-sense genomic RNA becomes the genome of the progeny viruses. The mRNAs are gene transcripts of the last third of the virus genome after the initial overlapping reading frame. These mRNAs are translated by the host's ribosomes into the structural proteins and a number of accessory proteins.[19] RNA translation occurs inside the endoplasmic reticulum. The viral structural proteins S, E, and M move along the secretory pathway into the Golgi intermediate compartment. There, the M proteins direct most protein-protein interactions required for assembly of viruses following its binding to the nucleocapsid.[21] Progeny viruses are then released from the host cell by exocytosis through secretory vesicles.[21]
Transmission
Human to human transmission of coronaviruses is primarily thought to occur among close contacts via respiratory droplets generated by sneezing and coughing.[22] The interaction of the coronavirus spike protein with its complement host cell receptor is central in determining the tissue tropism, infectivity, and species range of the virus.[23][24] The SARS coronavirus, for example, infects human cells by attaching to the angiotensin-converting enzyme 2 (ACE2) receptor.[25]
Taxonomy
The scientific name for coronavirus is Orthocoronavirinae or Coronavirinae.[2][3][4] Coronavirus belongs to the family of Coronaviridae.
- Genus: Alphacoronavirus
- Genus Betacoronavirus; type species: Murine coronavirus
- Species: Betacoronavirus 1, Human coronavirus HKU1, Murine coronavirus, Pipistrellus bat coronavirus HKU5, Rousettus bat coronavirus HKU9, Severe acute respiratory syndrome-related coronavirus, Severe acute respiratory syndrome coronavirus 2, Tylonycteris bat coronavirus HKU4, Middle East respiratory syndrome-related coronavirus, Human coronavirus OC43, Hedgehog coronavirus 1 (EriCoV)
- Genus Gammacoronavirus; type species: Infectious bronchitis virus
- Genus Deltacoronavirus; type species: Bulbul coronavirus HKU11
Evolution
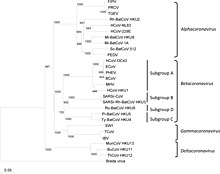
The most recent common ancestor(MRCA) of all coronaviruses has been placed at around 8000 BCE.[26] The MRCAs of the Alphacoronavirus line has been placed at about 2400 BCE, the Betacoronavirus line at 3300 BCE, the Gammacoronavirus line at 2800 BCE, and the Deltacoronavirus line at about 3000 BCE. It appears that bats and birds, as warm-blooded flying vertebrates, are ideal hosts for the coronavirus gene source (with bats for Alphacoronavirus and Betacoronavirus, and birds for Gammacoronavirus and Deltacoronavirus) to fuel coronavirus evolution and dissemination.[27]
Bovine coronavirus and canine respiratory coronaviruses diverged from a common ancestor in 1951.[28] Bovine coronavirus and human coronavirus OC43 diverged around the 1890s. Bovine coronavirus diverged from the equine coronavirus species at the end of the 18th century.[29]
The MRCA of human coronavirus OC43 has been dated to the 1950s.[30]
MERS-CoV, although related to several bat coronavirus species, appears to have diverged from these several centuries ago.[31] The human coronavirus NL63 and a bat coronavirus shared an MRCA 563–822 years ago.[32]
The most closely related bat coronavirus and SARS-CoV diverged in 1986.[33] A path of evolution of the SARS virus and keen relationship with bats have been proposed. The authors suggest that the coronaviruses have been coevolved with bats for a long time and the ancestors of SARS-CoV first infected the species of the genus Hipposideridae, subsequently spread to species of the Rhinolophidae and then to civets, and finally to humans.[34][35]
Alpaca coronavirus and human coronavirus 229E diverged before 1960.[36]
Coronaviruses vary significantly in risk factor. Some can kill more than 30% of those infected (such as MERS-CoV), and some are relatively harmless, such as the common cold.[19] Coronaviruses cause colds with major symptoms, such as fever, and sore throat from swollen adenoids, occurring primarily in the winter and early spring seasons.[37]Coronaviruses can cause pneumonia(either direct viral pneumonia or a secondary bacterial pneumonia) and bronchitis (either direct viral bronchitis or a secondary bacterial bronchitis).[38]The much publicized human coronavirus discovered in 2003, SARS-CoV, which causes severe acute respiratory syndrome (SARS), has a unique pathogenesis because it causes both upper and lower respiratory tract infections.[38]
Seven strains of human coronaviruses are known:
- Human coronavirus 229E (HCoV-229E)
- Human coronavirus OC43 (HCoV-OC43)
- Severe acute respiratory syndrome coronavirus (SARS-CoV)
- Human coronavirus NL63 (HCoV-NL63, New Haven coronavirus)
- Human coronavirus HKU1
- Middle East respiratory syndrome-related coronavirus (MERS-CoV), previously known as novel coronavirus 2012 and HCoV-EMC
- Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), previously known as 2019-nCoV or "novel coronavirus 2019"
The coronaviruses HCoV-229E, -NL63, -OC43, and -HKU1 continually circulate in the human population and cause respiratory infections in adults and children world-wide.[39]
[relevant? ]
Studies have shown that warm and wet air lowers both the influenza virus survival times and the virus to human transmission efficiency.[40] Airborne influenza virus survival may be be less at air temperatures above 30°C (86°F) where the humidity is above 50%, (exceeds an absolute humidity above 0.135 kg/m3).[41] Virus survival times were much lower at higher temperatures and higher relative humidity (e.g., 38°C [100°F], and relative humidity of >95%).[42]
Virus-laden droplet nuclei are more efficiently produced at lower relative humidity because of increased evaporation of expelled droplet particles, such that more virus remains airborne longer. In do survive and infectious in air with a low absolute humidity [such as <0.007 kg water/kg air].[43][44] When airborne droplets evaporate, then very light airborne droplet nuclei are formed. Virus in airborne droplet nuclei may remain in the air for many hours.[45]
[relevant? ]
The survival times at 80% RH and 40°C [104°F] were less than 7 hours for proxies of coronaviruses on stainless steel.*
The survival times at 50% RH and 40°C [104°F] were more than 24 hours.* There was a reduction in the ratio of virus to -3 Log10 (Nt/N0) in 24 hours.
The survival times at 20% RH and 40°C [104°F] were more than 120 hours.*There was a reduction in the ratio of virus to -3 Log10 (Nt/N0) in 120 hours.
The survival times at 50% RH and 20°C [68°F] were less than 7 days on stainless steel.*
* for proxies of coronaviruses on stainless steel. The proxies used were transmissible gastroenteritis virus (TGEV) and mouse hepatitis virus (MHV).[46][47]
Outbreaks of coronavirus types of relatively high mortality are as follows:
Severe acute respiratory syndrome (SARS)
In 2003, following the outbreak of severe acute respiratory syndrome (SARS) which had begun the prior year in Asia, and secondary cases elsewhere in the world, the World Health Organization(WHO) issued a press release stating that a novel coronavirus identified by a number of laboratories was the causative agent for SARS. The virus was officially named the SARS coronavirus (SARS-CoV). More than 8,000 people were infected, about ten percent of whom died.[25]
Middle East respiratory syndrome (MERS)
In September 2012, a new type of coronavirus was identified, initially called Novel Coronavirus 2012, and now officially named Middle East respiratory syndrome coronavirus (MERS-CoV).[53][54] The World Health Organization issued a global alert soon after.[55] The WHO update on 28 September 2012 said the virus did not seem to pass easily from person to person.[56] However, on 12 May 2013, a case of human-to-human transmission in France was confirmed by the French Ministry of Social Affairs and Health.[57]In addition, cases of human-to-human transmission were reported by the Ministry of Health in Tunisia. Two confirmed cases involved people who seemed to have caught the disease from their late father, who became ill after a visit to Qatar and Saudi Arabia. Despite this, it appears the virus had trouble spreading from human to human, as most individuals who are infected do not transmit the virus.[58] By 30 October 2013, there were 124 cases and 52 deaths in Saudi Arabia.[59]
After the Dutch Erasmus Medical Centresequenced the virus, the virus was given a new name, Human Coronavirus–Erasmus Medical Centre (HCoV-EMC). The final name for the virus is Middle East respiratory syndrome coronavirus (MERS-CoV). In May 2014, the only two United States cases of MERS-CoV infection were recorded, both occurring in healthcare workers who worked in Saudi Arabia and then travelled to the U.S. One was treated in Indiana and one in Florida. Both were hospitalized temporarily and then discharged.[60]
In May 2015, an outbreak of MERS-CoV occurred in the Republic of Korea, when a man who had traveled to the Middle East, visited 4 hospitals in the Seoul area to treat his illness. This caused one of the largest outbreaks of MERS-CoV outside the Middle East.[61] As of December 2019, 2,468 cases of MERS-CoV infection had been confirmed by laboratory tests, 851 of which were fatal, a mortality rate of approximately 34.5%.[62]



Comments
Post a Comment